In the laboratories of IIITH, a scientific breakthrough was taking shape. IIITH researchers, in collaboration with IHub-Data, were about to drastically change the landscape of drug discovery.
The Challenge: A Race Against Time
Developing a new medicine has always been a long, expensive, and high-risk journey—often taking over a decade and costing billions, with very few drug candidates making it to the finish line. Traditionally, researchers had to test thousands of compounds in the lab, a process that demanded huge effort and came with high failure rates, especially in the early stages. But things are changing. Computational protocols, especially models based on artificial intelligence and machine learning (AI/ML models), complement the tedious process in mitigating the time and cost exhaustion. At IIITH, researchers revolutionized the computational complementarity by creating datasets with advanced computer simulations powered by AI and machine learning to speed up drug discovery. While early models relied on static molecular data, they soon realized the real game-changer would be dynamic datasets that capture how molecules move and interact over time. That urgent need led to a breakthrough—one that could reshape the future of medicine.
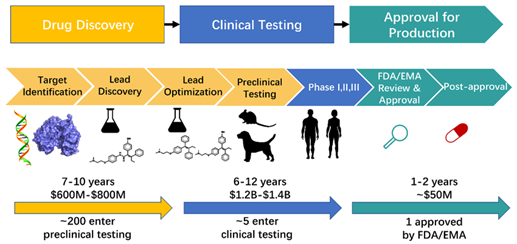
Figure 1: Reference – International Journal of Molecular Sciences (IJMS)
But this is changing.
Researchers at IIIT Hyderabad (IIITH), in collaboration with IHub-Data, are combining artificial intelligence (AI) and computer simulations to drastically speed up early-stage drug discovery. Instead of relying on static snapshots of molecules, they developed dynamic datasets that capture how these molecules move and interact over time—more closely mimicking what happens in the human body. This shift led to a breakthrough that could reshape the future of medicine.
A Breakthrough: The PLAS-20K Dataset
The team first launched PLAS-5K and later scaled it up to PLAS-20K —a massive, state-of-the-art dataset published in Nature Scientific Data, a leading open-access journal for high-impact datasets.
PLAS-20K includes nearly 20,000 detailed simulations of how small drug-like molecules bind to proteins in the body. It not only captures dynamic 3D coordinates but also includes calculated binding strengths—helping researchers train AI models to find promising drug candidates faster, more accurately, and at lower cost.
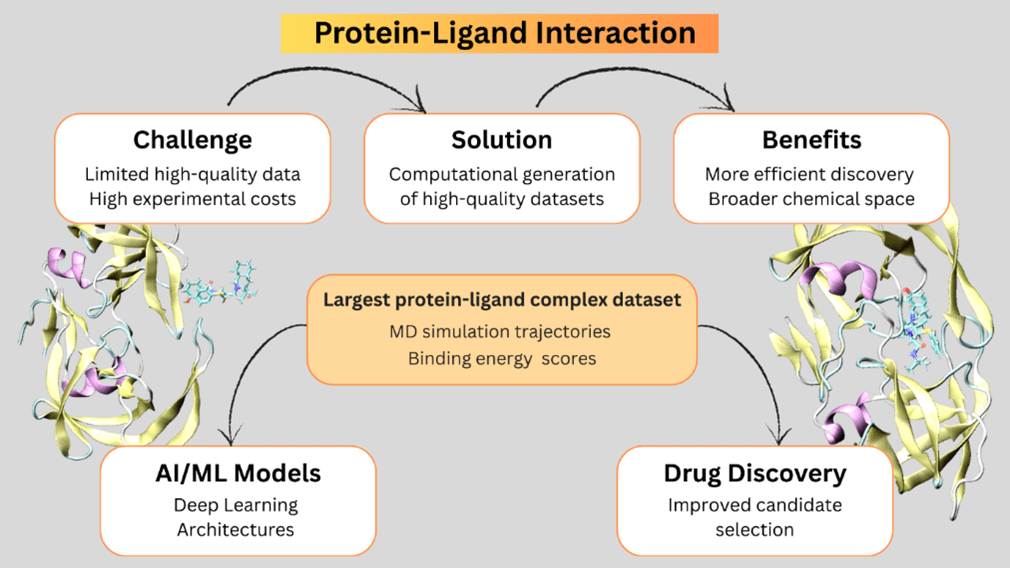
Figure 2: Protein-Ligand Interaction
Behind the Scenes: How PLAS-20K Was Built
Building PLAS-20K wasn’t easy. Each of the 20,000 protein-ligand complexes was carefully set up with care, simulated using molecular dynamics, and rigorously validated. One key step was calculating binding strength using a method called MM-PBSA and then comparing it to known experimental results. Interestingly, the models worked best when just two water molecules were included near the active site—subtle details like this helped improve the accuracy.
Inside the Dataset: Two Powerful Variants
The PLAS-20K dataset comes in two variations, both available at India Data:
Variation 1: Offers cleaned-up 3D structures of protein-drug interactions in near-physiological conditions. It includes averaged binding strengths and helps researchers understand how drugs behave inside the body.
Variation 2: Goes further by capturing how interactions change over time. This dynamic view includes snapshots across simulation timelines and energy-related data—critical for training next-generation AI models.
Dataset Usage So Far:
Testing the Dataset: Training AI for Binding Predictions
To test how useful PLAS-20K really is, the team trained AI models like OnionNet, using the dataset’s 3D coordinates and binding energy values. Through a testing method called 10-fold cross-validation, they found that these AI models performed reliably well.
Importantly, adding dynamic interaction data improved the model’s ability to predict whether a molecule would bind strongly to a protein. This opens exciting possibilities: better predictions, fewer failed drugs, and much faster discovery timelines.
What’s next? The team is expanding PLAS-20K by also including “negative examples”—cases where molecules don’t bind well. This helps AI learn to tell the difference between real drug candidates and misleading ones.
PLAS-20K: A New Era for Drug Discovery
PLAS-20K isn’t just a dataset. It’s a platform for transforming how we discover and develop medicines.
By integrating high-quality molecular simulations with AI-ready formats, this resource makes it easier for scientists to test ideas, filter out failures early, and bring life-saving treatments to market faster. As the role of AI in biology grows, datasets like PLAS-20K will become the backbone of a new, smarter approach to drug discovery.
To explore/download the dataset and learn more about our Drug Discovery R&D, please visit:
Author
Prof. Deva Priyakumar – Professor, Ph.D (Pondicherry University)
Email: deva@iiit.ac.in
Website URL: Deva Priyakumar U – IIIT Hyderabad
Dr. Prathit Chatterjee, IIITH
LinkedIn URL: Prathit Chatterjee
Publisher
Sangita Rathod, IHUB-Data
LinkedIn URL: Sangita Rathod
